New technique scans drop of blood for all human viruses across time
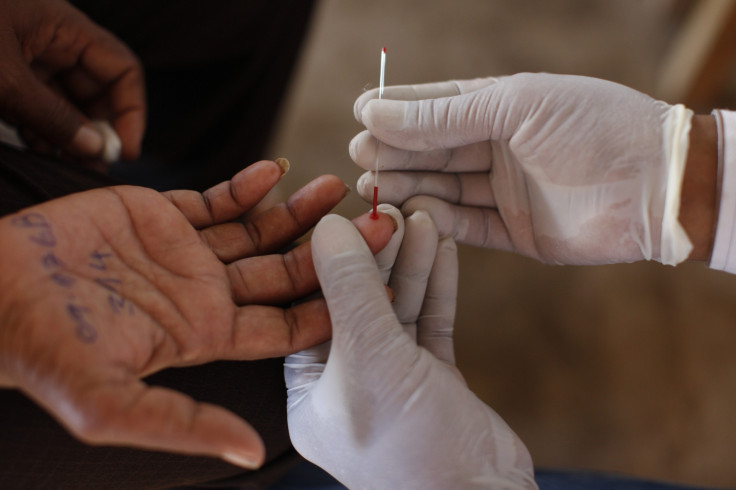
All it takes is a drop of blood to detect all existing and past virus infections using a technique developed by Howard Hughes Medical Institute (HHMI) researchers.
Called VirScan, the method can run the whole gamut of human viruses in the drop to check against the person's health in a single test. It does this by screening the blood for antibodies against any of the 206 species of viruses known to infect humans.
The tests can be performed for about $25 (£16) per blood sample.
Besides being more efficient than current diagnostics that need to scan for viruses in successive tests, VirScan expands opportunities to study viral infections in large populations.
Stephen Elledge, an HHMI investigator at Brigham and Women's Hospital, who led the development of VirScan calls it "one-stop shopping."
Elledge and his colleagues used VirScan to screen the blood of 569 people in the United States, South Africa, Thailand, and Peru, examining about 100 million potential antibody/epitope interactions. They found that on average, each person had antibodies to ten different species of viruses.
The researchers also found that people infected with HIV had antibodies against many more viruses than did people without HIV.
Elledge says the team was surprised to find that antibody responses against specific viruses were surprisingly similar between individuals, with different people's antibodies recognising identical amino acids in the viral peptides.
Tested on patients known to be infected with particular viruses, including HIV and hepatitis C the VirScan analysis proved it worked.
"We were in the sensitivity range of 95 to 100 percent for those, and the specificity was good -- we didn't falsely identify people who were negative," says Elledge.
The findings are reported in the June 5, 2015 issue of the journal Science.
Antibody-epitope interaction
The immune system produces pathogen-specific antibodies in large numbers when it encounters a virus for the first time, and it can continue to produce those antibodies for years or decades after it clears an infection.
This is how VirScan not only identifies viral infections that the immune system is actively fighting, but also provides a history of an individual's past infections.
The team synthesised more than 93,000 short pieces of DNA encoding different segments of viral proteins. They introduced those pieces of DNA into bacterial viruses called bacteriophage.
Each bacteriophage manufactured one of the protein segments -- known as a peptide -- and displayed the peptide on its surface. As a group, the bacteriophage displayed all of the protein sequences found in the more than 1,000 known strains of human viruses.
To perform the VirScan analysis, all of the peptide-displaying bacteriophage were allowed to mingle with a blood sample. Antiviral antibodies in the blood find and bind to their target epitopes (embedded in proteins on virus surface) within the displayed peptides.
These antibodies are retrieved and washed to remove all but the bacteriophage that cling to them.
The DNA sequencing of the bacteriophages can identify the viral protein pieces picked by antibodies in the blood sample to identify the viruses.
It takes about 2-3 days to process 100 samples at present but the team expects to improve on this soon.
© Copyright IBTimes 2025. All rights reserved.




















