Can the woolly mammoth play a role in saving rhinos from extinction?
New technologies applied on ancient bones can provide vital knowledge about genetic threats.
Let's be clear. Saving rhinoceros species from extinction will mainly be down to hard practical conservation management, such as anti-poaching efforts, habitat protection and combating illegal trade in rhino horn.
Thus, even though emerging technologies in genomics, bioengineering and reproductive biology may provide valuable knowledge and novel approaches, it is important that these new technologies do not lead to funding being diverted from the practical conservation work.
Nonetheless, a recent revolution in DNA technology now opens up for cost-effective and large-scale mapping of rhinoceros biodiversity at its most fundamental level – and this could lead to several important insights that can help conservation decision-making.
In several rhino species, the number of individuals that remain is now so small that inbreeding and low genetic diversity may constitute a hindrance to the recovery of these species. I write 'may', because we simply do not yet know whether such genetic threats are having a real impact on the current health of rhino populations.
Theory predicts that small populations will be affected by inbreeding and loss of genetic diversity. As a result, harmful mutations can accumulate in the genome and populations will lose the capacity to adapt to changes in the environment.
But until recently, it has been very difficult to investigate whether such genetic problems exist in wild rhino species. Estimating inbreeding levels used to require detailed pedigrees, something that is nearly impossible to obtain for wild and elusive species. Moreover, genetic mutations that are harmful are only expected to occur at a few places in an individual's genome, and discovering these few mutations in a genetic code that consists of up towards the billion DNA letters would have been very difficult and expensive only a few years ago.
Today, however, modern DNA technology makes it possible to sequence a complete genome with high accuracy in a matter of days, and at a cost of around €500 (£430) per individual. This means that it now is relatively easy to estimate inbreeding levels and genetic changes that may have a negative impact on the health of individual animals.
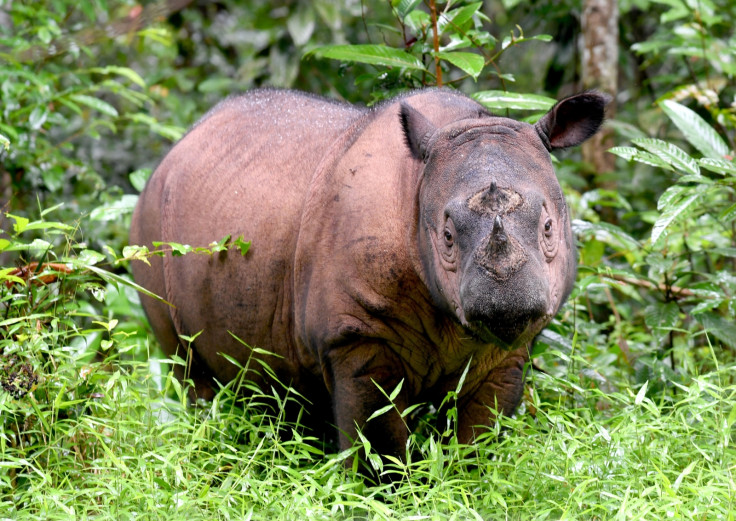

Together with colleagues from several institutes around the world, we have recently initiated a genome project on the Sumatran rhino, with the aim to better understand the genetic threats facing this species and to aid current conservation management. Although historically abundant and widespread across much of south-east Asia, the Sumatran rhino is nowadays one of the most threatened mammals in the world. Only some 100 individuals remain, in isolated populations on the islands of Sumatra and Borneo.
To estimate the impact of this decline, we are using a form of 'genomic time-travel' to look at genetic diversity in rhinos that lived in historical times. We do this through sequencing a large number of genomes from museum samples that are about 100 years old, and compare these with genomes from modern-day samples. This will allow us to quantify the extent of genomic erosion that has taken place as a consequence of the decline in Sumatran rhino populations. Also, using the 100-year-old museum specimens as a baseline will enable us to identify any harmful genetic mutations that have increased in frequency in recent years.
So how can such a temporal approach be used to help in saving the Sumatran rhino, and other species, from extinction? First of all, these analyses will make it easier for decision-makers to understand the scale of the genetic threat, and thus to what extent conservation actions are needed to lessen the effect of inbreeding, harmful mutations and loss of genetic diversity.

One example of such an action could be to move individuals from one population to the other, since this would alleviate inbreeding problems and introduce new genetic variation. A comparison of genomic divergence among populations will also help conservation managers assess how likely it is that such a translocation would succeed. Finally, sequencing the genomes from living individuals and identifying inbreeding levels as well as which harmful mutations they carry can be used to select individuals for captive breeding as well as possible translocation programmes.
But can genetic problems really lead to a species going extinct? The simple answer is that we don't know. Theoretical work suggests that genetic problems can become so severe that population growth becomes negative, which eventually would cause extinction. There might even exist thresholds of genomic erosion, or points of no return, from which species are unable to recover.
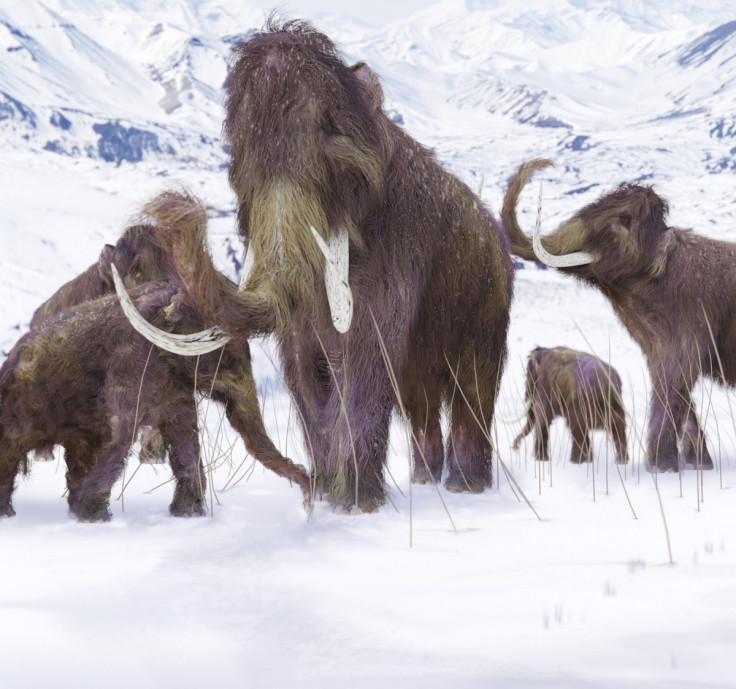
One way to test this theory is to investigate whether, and to what extent, species that are already extinct went through genomic erosion just before they disappeared. We are now examining this in several extinct species. So far, we have generated genomes from both the woolly rhinoceros and the woolly mammoth, by extracting and sequencing DNA from bones and teeth that are several thousands of years old.
At least in the woolly mammoth, where we have compared genomes from different time points during its decline towards oblivion, it appears that genetic factors may have played a role in the extinction. This illustrates the importance of establishing how far modern-day endangered species have travelled along the road of genomic erosion, and whether they have already crossed the point of no return.
Love Dalén is Professor of Evolutionary Genetics at the Department of Bioinformatics and Genetics, Swedish Museum of Natural History
© Copyright IBTimes 2025. All rights reserved.