Our ancestors defeated a virus from the same family as HIV 11 million years ago
Scientists studied ''viral fossils'' to find out how how the HERV-T retrovirus went extinct.
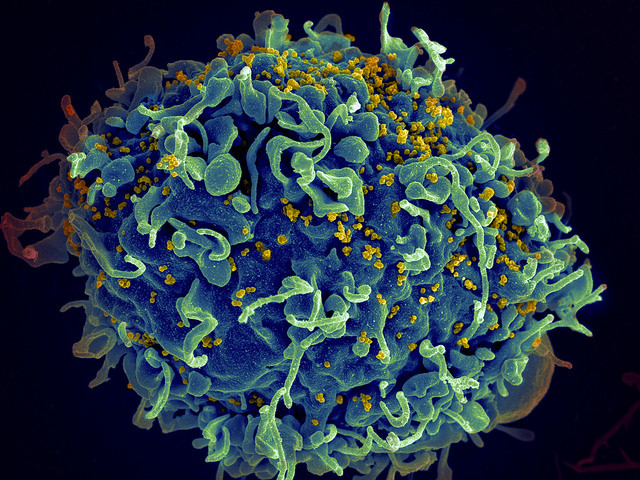
Viral fossils have revealed the secret of how our ancestors defeated an ancient retrovirus known as about 11 million years ago. Ancient hominids might have ''hijacked'' one of the virus' gene to prevent it from infecting their cells.
Studying the evolution of animals over million of years has been possible thanks to the many fossils of their extinct relatives that remained, spread out across the world. However, investigating the long history of viruses has proven a lot more challenging since they typically do not leave a trace of their existence.
The only exception are retroviruses, the virus family that HERV-T but also HIV belong to. Scientists are able to study the evolution of these retroviruses, because when they infect an animal, their genetic material is integrated to the genome of the infected cells.
When these cells are precursors of sperm or egg cells, then the genes of the virus can be passed on to the animal's offspring, ultimately leaving genetic traces that can still be observed today in modern animals.
Scientists consider this to be a genetic ''fossil record'' of extinct viruses.
In a study now published in the journal eLife, researchers led by Paul Bieniasz from the Rockefeller University studied the genetic fossil record of HERV-T. This viral fossil evidence indicated that HERV-T began to replicate in primates some 40 million years ago.
However, the scientists also found that a specific viral gene was later hijacked 13 to 19 million years ago leading to the extinction of HERV-T in hominids 11 million years ago.
"Unfortunately we can't be very precise in our time estimates. However, it is quite likely that the hijacked viral gene had antiviral activity immediately upon incorporation into the genome of an individual host. It would then have taken some unknown number of generations for that gene to spread to all members of the ancestral hominid species", Bieniasz told IBTimes UK.
Hijacking the viral gene
In this study, the scientists compiled and analysed a large catalogue of HERV-T fossils in old-world monkey and great ape genomes (including human genomes). They used this information to recreate an outer envelope protein of the ancient retrovirus, which allowed it to bind to hosts' cells and to infect them.
Experiments suggested that this viral protein interacted with a protein called MCT1 on the surface of the cells. It is this interaction that led to the cells becoming infected.
Looking at the genetic ''fossil record'' of HERV-T, the scientists found that one gene in particular was well preserved in the genome of humans. The scientists believe that ancient hominids might have ''hijacked'' this viral gene and used the protein it produced to remove the MCT1 protein from the surface of their own cells.
In the absence of MCT1, the retrovirus would not have been able to infect the cells and it would have ended up becoming extinct.
Weapon against HIV?
These findings provide an interesting insight into the evolution of retroviruses. However, they do not offer concrete solutions as to how we can fight current retroviruses like HIV.
"HIV-1 is a quite different virus, and uses different receptor molecules to enter cells. It is true that the general idea of blocking receptors to interfere with virus infection is plausible – there are drugs (licensed and under investigation) to treat and prevent HIV-1 infection that work in this way. But we didn't need to know about HERV-T extinction to do this," Bieniasz said.
Still, the scientists note that this work might have another potential, entirely unexpected application.
"It turns out that the normal cellular function of MCT-1, the receptor used by HERV-T millions of years ago to infect cells, is to transport metabolites across cell membranes. Certain types of tumours are highly dependent on these metabolites, and probably as a consequence, make large amounts of MCT-1 on their surface," Bieniasz explained.
They are now beginning a new project to use the HERV-T envelope protein they have recreated in this study to mark, and possibly kill, tumour cells that over-express MCT-1.
© Copyright IBTimes 2025. All rights reserved.




















