Swine flu: 2009 pandemic that killed 17,000 came from one small region in Central Mexico
Origins of the H1N1 influenza pandemic have at last been determined in detail.
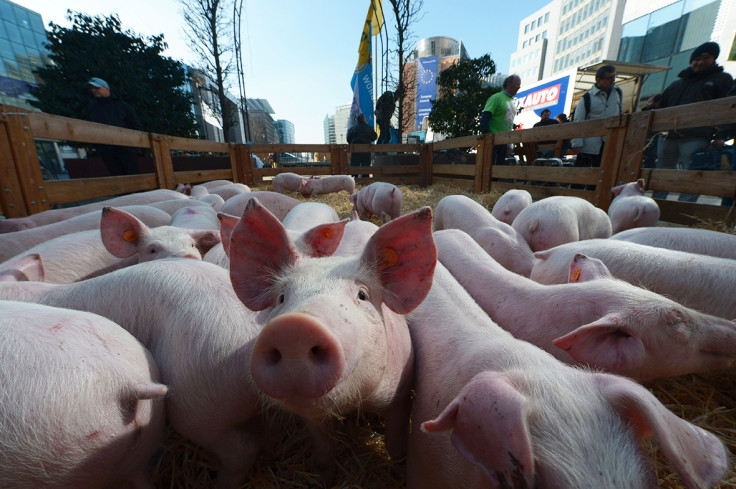

The H1N1 swine flu pandemic of 2009 originated from pigs in a small region in Central Mexico, scientists have discovered. It is the first time that the geographic origin of an influenza pandemic has been determined in such specific detail.
The disease spread quickly from pigs to humans and from the Central American country to the rest of the world, killing more than 17,000 people.
The most closely related ancestor to the H1N1 virus were identified in Asian swine, and it wasn't previously clear where the strain came from.
For the study, published in eLife, the researchers conducted a large genetic analysis of different Mexican pigs' virus strains in order to identify the precise location and the main molecular transformations that allowed the 2009 pig influenza virus to jump to humans.
58 genomic sequences
The scientists, from the National Institute of Health in the US and Avi-Mex, a Mexican veterinarian company, looked at 58 whole-genome sequences from influenza viruses collected from Mexican pigs.
The researchers identified viral sequences previously not known to exist in Mexico, or in any other part of the world. Flu viruses are constituted of eight mini-chromosomes and can exchange genetic segments with other strains when both infect the same cell – a process called reassortment. All the 58 genomic sequences analysed here had reassortments not seen elsewhere.
These findings indicate that the 2009 H1N1 was in fact a derivative of two different strains of swine influenza – one that had previously been circulating in Europe and Asia and another that was circulating in the Americas, especially North America.
The scientists are also able to say that a 'parent virus' of this hybrid strain was present for 10 years in Mexican pigs before it became able to transfer to humans, which happened in 2009.
"This virus came from, and was confined to, a very small geographic area and had been there 10 years before one emergent strain gained the capacity to infect humans," author Adolfo García-Sastre says. The area of Central Mexico in question had not previously been considered a pandemic risk.
Surveillance of future viruses
The researchers hope their work will help better spot and prevent transmissions of viruses from animals to humans. "Knowing where and how an animal influenza virus infects humans and spreads all over the world helps us understand how we can reduce risk of these pandemics," says García-Sastre.
By solving the origins and the reassortments that led to the 2009 pandemic, researchers now believe they can identify existing "brothers and sisters" of the virus to further study the type of mutations needed to allow a virus to jump into humans.
It is also crucial to study worldwide surveillance of active flu in pigs to make sure that reassortments leading to human infection do not happen again.
"We need to monitor the viruses that are circulating, and try to stop mixing influenza strains from different geographic locations," he says. "This study also shows that you cannot ignore small geographic areas with pig farms – places in which the 2009 pandemic originated and which the next, perhaps more severe global flu, may come from", concluded García-Sastre.
© Copyright IBTimes 2025. All rights reserved.